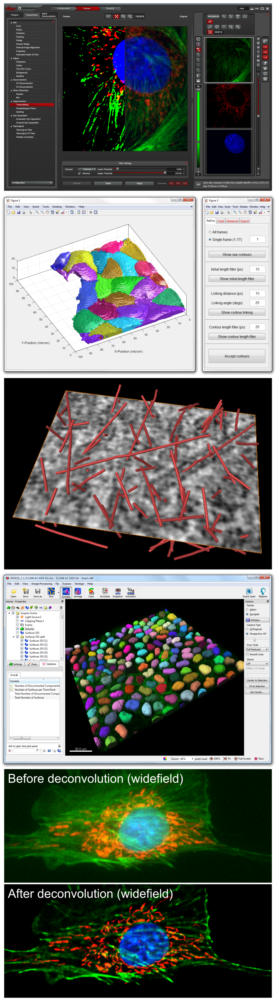
Software
A variety of open-source and proprietary image processing and analysis software is supported within the facility.
-
Amira
Designed for segmentation, visualisation and quantification of 3D and 4D imaging data, in particular, electron microscopy data. The version available in the facility includes specific packages for life science image processing and analysis.
-
Arivis Vision4D
For visualisation, processing, and analysis of massive (10s to 100s of GB), multidimensional images. Vision4D has capability for volumetric and surface renders of images/movies. Can be used to segment objects and measure shape descriptors (including volume, surface area and ellipticity).
-
CellProfiler
Pipeline-based image analysis software specialising in object segmentation and measurement. It incorporates many advanced algorithms for accurate segmentation and processing of objects of interest.
-
FLIMFit
Analysis software for fluorescence lifetime imaging (FLIM) experiments. It allows the fitting of multiexponential models to time resolved data, and can incorporate image segmentation. The efficient fitting algorithm allows global analysis of large data sets to be performed in under a minute.
-
Huygens
For image restoration via deconvolution. Deconvolution is an algorithm-based approach for reassigning out-of-focus light to its origin, thus improving signal-to-noise in images. This is most evident for images collected on widefield systems; however, it can also be applied to confocal, STED and lightsheet images. Huygens has additional functionality for standard image processing and analysis operations.
-
ImageJ/Fiji
A highly flexible freeware image processing and analysis tool from NIH. Available as standard ImageJ or the advanced Fiji version, which includes additional commonly used plugins. Functionality of either package can be extended further using a vast online library of peer-reviewed plugins.
-
Imaris
For segmentation, visualisation and quantitation of 3D and 4D imaging data, which is specialised for processing images of life sciences samples. It includes high-quality rendering of complex multi-stage animations, and the functionality can be extended via use of custom ImageJ/Fiji macros or MATLAB scripts.
-
Leica LAS-X
A commercial software platform for microscope control and for image acquisition, management, processing and analysis. This package (or equivalent) is installed on all facility Leica systems. An additional core functionality version is freely available online for image viewing and basic processing and analysis.
-
MATLAB
A versatile scripting-based programming language, widely used for image processing and analysis due to its emphasis on efficient processing of numerical arrays. It is ideal for creating custom analysis pipelines or developing new algorithms, and it also has the capability for compiling programs into standalone executables with user-friendly interfaces.
-
MIA: Modular Image Analysis
A Fiji plugin which provides a modular framework for assembling image and object analysis workflows. Detected objects can be transformed, filtered, measured and related. Analysis workflows are batch-enabled by default, allowing easy processing of high-content datasets. MIA is developed in-house at the Wolfson Bioimaging Facility, in response to the needs of our users.
-
PicoQuant SymPhoTime64
Analysis software for fluorescence lifetime imaging (FLIM) and fluorescence correlation spectroscopy (FCS) data allows the fitting of multiexponential models to time resolved data on a pixelwise basis.
-
Volocity
A general purpose image processing and analysis suite specialising in visualisation of 3D and 4D data.