Scientists have previously designed protein catalysts from scratch using computers, but these are much less capable than natural enzymes. To improve their performance, a technique called laboratory evolution can be used, which American chemical engineer Frances Arnold pioneered and for which she received the Nobel Prize in 2018. Directed evolution imitates natural selection, allowing scientists to use the power of biology to improve the ability of proteins to carry out tasks such as catalysing a specific chemical reaction.
But although the research team had recently used laboratory evolution to improve a designed enzyme by more than 1,000 fold, it was unknown how evolution boosts its activity. Until now.
Lead author Professor Adrian Mulholland of Bristol's School of Chemistry said: "Evolution can make catalysts much more active. The thing is, evolution works in mysterious ways: for example, mutations that apparently improve catalysis often involve changes in amino acids far from the active site where the reaction happens.”
"We wanted to understand how evolution can transform inefficient designer biocatalysts into highly active enzymes," the first author of the study Dr Adrian Bunzel, said.
To do so, the international research team from Bristol, the ETH Zurich and the University of Waikato (NZ) turned to molecular computer simulations. He added: "These show that evolution changes the way the protein moves – its dynamics. Put simply, evolution ‘tunes’ the flexibility of the whole protein."
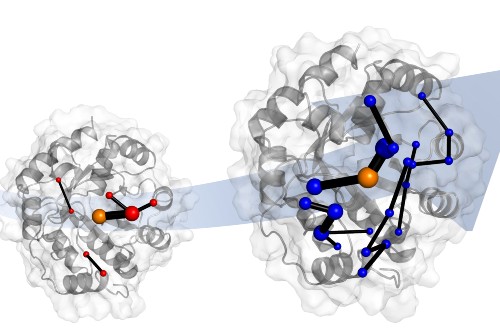
The team also identified the network of amino acids in the protein responsible for this ‘tuning’. These networks involve parts of the protein that are changed by evolution.
Dr Bunzel explained: "After evolution, the whole protein seems to work together to accelerate the reaction. This is important because when we design enzymes, we often focus only on the active site only, and forget about the rest of the protein."
Professor Mulholland said: "This sort of analysis could help to design more effective ‘de novo’ enzymes, for reactions that previously we could not target."
The research, published in Nature Chemistry, reveals how evolution makes designer enzymes more powerful, paving the way to tailor-made catalysts for green chemistry.
The researchers will now use their findings to help design new protein catalysts.
Paper
'Evolution of dynamical networks enhances catalysis in a designer enzyme' in Nature Chemistry by HA Bunzel, JLR Anderson, D Hilvert, VL Arcus, MW van der Kamp and AJ Mulholland